The Promise and Perils of Machine Learning in Pharmaceutical Drug Development

A look at GlaxoSmithKline's efforts to incorporate machine learning into their drug development processes.
Pharmaceutical drug development has long been an expensive process, in part due to high attrition rates for candidate drug products. Enter machine learning – but, not all aspects of drug product development are equally suitable for machine learning to address. Among the big pharmaceutical firms, GlaxoSmithKline (GSK) provides a good case study of how machine learning is being incorporated to benefit early drug development, and where the pitfalls may lie.
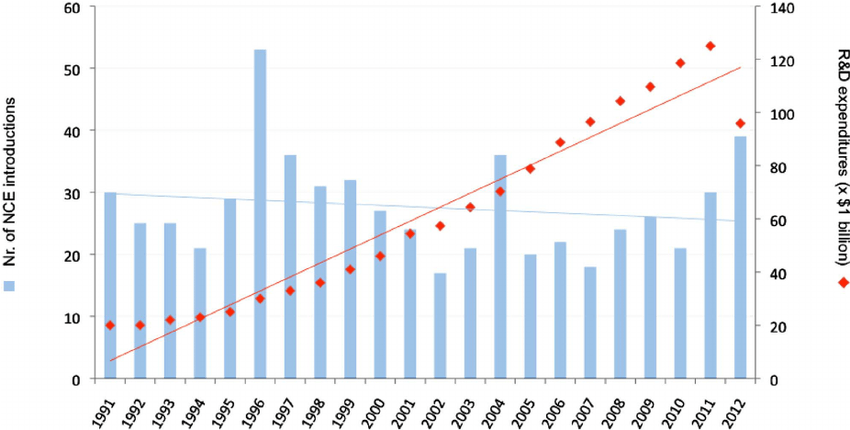
Machine learning has the potential to strengthen GSK’s competitive advantage in R&D. The pharmaceutical industry has seen a decline in R&D productivity over the last decades, with increasing R&D spend but a relatively flat number of FDA approvals per year, as shown in the chart below [1].
One of the main reasons behind the declining trend is the increased rate of failure of drug development programs [2]. Consequently, it’s critical to GSK’s competitive advantage to limit attrition in its pipeline of drug candidates – and that begins in early R&D, with tasks such as understanding disease pathology, identifying a target and designing a molecule to hit it. GSK’s researchers have made strides towards employing machine learning to support these tasks, for instance using a semi-supervised approach on gene-disease association data in order to predict potential new drug targets [3]. They are also seeking to apply machine learning to analyzing normal vs. tumor-derived genomic, metabolomic, proteomic, transcriptomic and other kinds of -omics data in order to characterize the pathology of cancer, in hopes of finding novel cancer targets [4].
In addition to encouraging internal integration of machine learning into drug product development processes, GSK’s management has focused on partnerships with AI firms. Both of these initiatives are ways for GSK to start exploring the potential of machine learning in the next couple of years, with ramifications that will extend beyond that into the next decade. To help centralize the push into machine learning and related technologies, GSK appointed their first Chief Digital Officer, Karenann Terrell, in 2017 [5]. Around the same time, GSK entered into a collaboration with Exscientia, an AI drug design company, seeking to use AI to design molecules for up to 10 GSK-selected disease targets [6]. Along similar lines, GSK in 2018 entered into a collaboration with Cloud Pharmaceuticals also to design tailored molecules for GSK-selected targets [7]. The two partnerships center around a task that machine learning is well suited for: predicting features of a drug molecule that will make it the best fit for a target, given a large set of previous molecules and targets that matched well together. Thus far, GSK management has done well in applying machine learning to prediction problems (to their benefit), rather than problems centered on causal explanation (which could have been a pitfall).
Looking ahead, there are further actions that GSK management could take in the short and medium terms to fully benefit from the opportunities provided by machine learning technologies. These come in three main categories: (1) acquiring/licensing more data to work with, (2) finding additional areas that machine learning can be useful in, and (3) evaluating the outcomes of initiatives. When looking for more data, GSK might consider investing in a portfolio of many different kinds of sources, including for instance licensing in EMR/claims data or sponsoring new academic initiatives to collect genomic or proteomic data. When considering additional areas in which machine learning can be applied, GSK must continue to focus on prediction, and avoid issues that center on an explanation of why a result happens. One possible area is the prediction of adverse events from early-stage molecule data, early enough where the design of the molecule might still be modified to alleviate the effect, making it less likely to attrition out during clinical trial testing. Another is prediction of drug response biomarkers, such that GSK can seek out patients more likely to respond well to their drug in clinical trials, raising the likelihood of success and thus making it more likely they can reach FDA approval, at least for treating that patient subset. And finally, GSK needs to create metrics (such as evaluating improvement in attrition rate from preclinical to Phase I) to decide which machine learning initiatives they should focus their resources on.
Although GSK is set to reap significant benefits to its drug product development from machine learning, there are still unresolved issues, namely that it is a “black box”, and no substitute for scientific expertise in explaining the underlying biological mechanism behind why a drug could work. The ability to elucidate the mechanism is also important when filing for approval with the FDA. With that in mind, future discussion might center around the question of, “How can we address the issue of low transparency into why machine learning made a particular prediction?” whether that be for drug targets, biomarkers, or more. (798 words)
Footnotes
[1] Kenneth Fernald, T.C. Weenen, Kelsey Sibley, and Eric Claassen, “Limits of Biotechnological Innovation,” Technology and Investment 43020(4) (2013): 168-178, [https://www.researchgate.net/publication/266175031_Limits_of_Biotechnological_Innovation], accessed November 2018.
[2] Bruce Booth and Rodney Zemmel, “Prospects for productivity,” Nature Reviews Drug Discovery 3 (2004): 451-456, [https://www.nature.com/articles/nrd1384], accessed November 2018.
[3] E Ferrero, I Dunham, and P Sanseau, “In silico prediction of novel therapeutic targets using gene-disease association data,” Journal of Translational Medicine 15(1) (2017): 182, [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5576250/], accessed November 2018.
[4] Silvana Valdebenito, Emil Lou, John Baldoni, George Okafo, and Eliseo Eugenin, “The Novel Roles of Connexin Channels and Tunneling Nanotubes in Cancer Pathogenesis,” International Journal of Molecular Sciences 19(5) (2018): 1270, [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5983846/], accessed November 2018.
[5] “Karenann Terrell appointed Chief Digital & Technology Officer, GSK,” press release, July 25, 2017, on GSK website, [https://www.gsk.com/en-gb/media/press-releases/karenann-terrell-appointed-chief-digital-technology-officer-gsk/], accessed November 2018.
[6] “Exscientia Enters Strategic Drug Discovery Collaboration with GSK: Pre-clinical collaboration focused on up to 10 targets nominated by GSK,” press release, July 2, 2017, PR Newswire, [https://www.prnewswire.com/news-releases/exscientia-enters-strategic-drug-discovery-collaboration-with-gsk-632029553.html], accessed November 2018.
[7] “Cloud Pharmaceuticals forms Drug Design Collaboration with GSK,” press release, May 30, 2018, Business Wire, [https://www.businesswire.com/news/home/20180530006184/en/Cloud-Pharmaceuticals-forms-Drug-Design-Collaboration-GSK], accessed November 2018.



While IBM Watson Health was concerned with making the functions of AI transparent to garner trust, its interesting that GSK needs to understand why the AI comes to certain results in order to back into experiments such that they can explain why drugs work when applying for FDA approval. It seems that, in order to keep the funnel of possible solutions open, GSK should ensure that there are not biases in how we understand nature today that would eliminate solutions in ways that we do not yet understand. Additionally, there may be a need for a parallel algorithm that can test possible experiments using “successful” compounds from their current AI in order to more quickly arrive at scientific explanations.
It is very interesting to think shortcomings in understanding the “why” aspect of biochemical and pharmaceutical processes. There is a whole field of researchers devoted to creating better and more accurate models of how molecules interact with each other, but I wonder whether ML could be applied to the understood chemical reactions and processes to create a good enough model at the cellular level. I am picturing an AI program that is given biological and chemical inputs, and scored based on how close to the outputs it gets, using a base data set of known chemical, biological and pharmaceutical reactions and their products.
Thanks for sharing this article. We’ve often discussed the notion that AI is not replacing human expertise but rather being used as a supplement to that human expertise. I think using machine learning early in the drug development process to help address attrition is fantastic in theory but it definitely raises some concerns. My main concerns around this have to do with biases. Would the inputs to this ML algorithm further proliferate existing biases and unknowns thus limiting a portion of the funnel of possible breakthrough ideas in the field of medicine? I understand the costliness of drug development and the sunk costs of attrition in this process which is why I think this is good idea, however, I think there needs to be a way to supervise, monitor and check against some of the unknowns and biases that might be stunting the effectiveness of the AI.
One concern I have with the use of ML to identify candidate drugs is that it risks emphasizing incremental progress over revolutionary new treatments. AI algorithms are excellent at predicting future outcomes based on past results. Because of this, I would guess that most of the drugs recommended by an algorithm would be inspired by drugs that have worked before. Would that reduce the likelihood of unexpected and serendipitous breakthroughs in medicine, where we discover a drug unlike any other we’ve had before?